Rosetta DARC Docking
This protocol follows the DARC docking procedure.
It can take as input either an AtomStruct (where you would need to define the residue(s) where the docking will be
performed around) or a SetOfStructROIs (to directly define the region as a Structural Regions Of Interest).
By default, DARC works only matching the shape of the ligand and receptor, but the user can input a electrostatic grid built by AutoDock to take into account also electrostatics, which might substantially increase the performance.
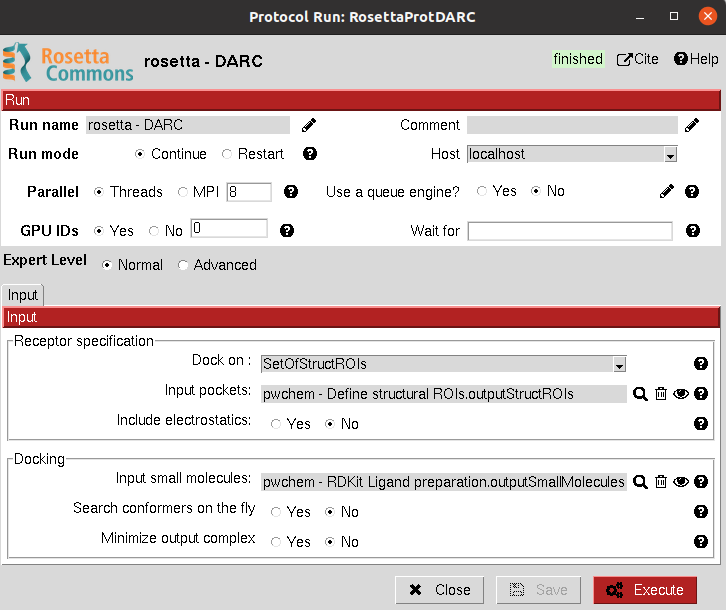
Input
Note
All parameters include a help button that gives further information for each of them.
The results of these protocols are a SetOfSmallMolecules, containing the predicted binding poses for the input
molecules. The user can visualize them using Analyze Results, which will display the General SmallMolecules viewer.
Using this viewer you can also visualize the rays that define the “pockets” where the molecules are docked.
Test
This protocol has an integrated test that can be run using the following command:
scipion3 tests rosetta.tests.test_darc.TestDARC